Did our cells evolve out of a slow, symbiotic dance?
Loading...
"Once upon a time, there was one cell which gave rise to all life on Earth," Buzz Baum, a cell biologist at the University College London, begins the story of life on our planet. "That's clear."
But the details of the next big step are far from clear.
Some simple single-celled organisms, prokaryotes, somehow evolved into more complex organisms, eukaryotes, about 2 billion years ago. But scientists have only hints about that key step in cellular evolution.
The story has largely been speculative, based on DNA analysis of organisms alive today. But last year a team of scientists discovered a prokaryote microbe that is remarkably closely related to eukaryotes.
The discovery of Lokiarchaeum, named for a hydrothermal vent called Loki's Castle near where the organism was found, deep in the Arctic Ocean, might help scientists clarify those speculative models.
Dr. Baum and two of his colleagues discuss those changes in an opinion paper published Thursday in the journal Trends in Cell Biology. They propose that the first eukaryotes arose out of a "long, slow dance between kingdoms," as study co-author Mukund Thattai of the National Centre for Biological Sciences in India said in a press release.
Life on Earth today belongs to two groups: prokaryotes and eukaryotes. Eukaryotes are both single- and multi-cellular organisms, encompassing everything from plants to animals, hummingbirds to humans, and fish to fungi. Prokaryotes are all single-celled organisms that fall into two different categories: bacteria and archaea.
Eukaryotic cells are more complex than prokaryotes. A eukaryotic cell has organelles, like mitochondria, that prokaryotes don't have, and a nucleus, while prokaryotes can be thought of as essentially balloons of DNA and protein.
So how did one emerge out of the other?
Over years of research, several models have been proposed, then subsequently rewritten or discarded, as new data has been collected, says Joel Dacks, Canada Research Chair in evolutionary cell biology and an associate professor at the University of Alberta, who was not part of the research team. He tells the Monitor, "I would say, right now there isn't a strongly favored model."
One model proposed by scientists suggested that an ancestral single-celled organism absorbed another prokaryote, putting it to work, and eventually making it an organelle: the mitochondria. That explanation made sense and has been largely accepted, as the mitochondria has its own DNA, distinct from the rest of the DNA in a eukaryote.
Some scientists previously suggested that such phagocytosis, as the cell-cannibalism is called, could explain how eukaryotes evolved from prokaryotes, says Gautam Dey, an evolutionary cell biologist at the University College London. But, he tells the Monitor, "This would have been quite a fast process if this is how things really went down."
Dr. Dey, Dr. Thattai, and Baum propose a more gradual process.
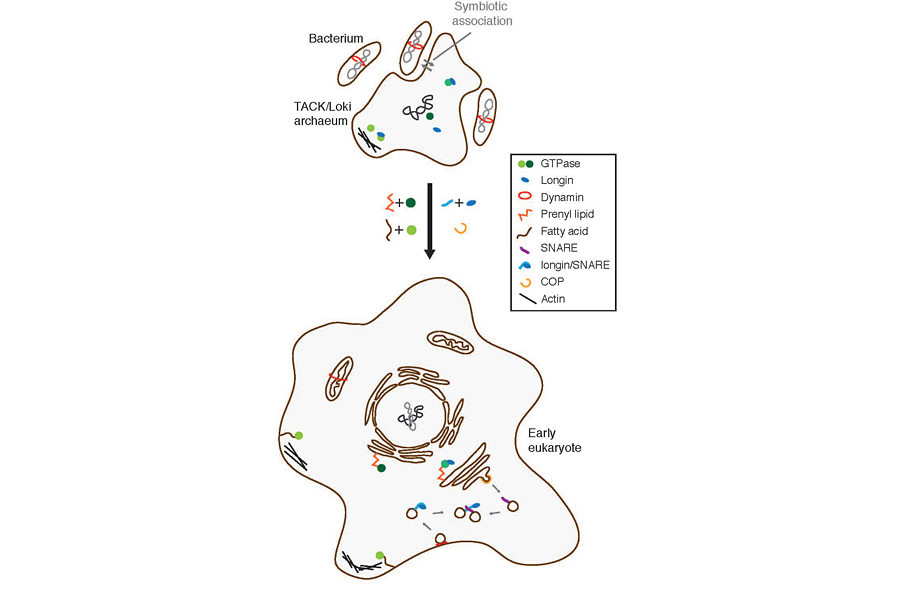
They suggest that an archaeon and a bacterium had a symbiotic relationship, exchanging material over time, gradually becoming closer and closer. As the archaeon incorporated more and more material from the bacterium within its own genome, it slowly evolved into the first eukaryote. And this would have happened over timescales of hundreds of thousands, maybe even millions, of years.
In other words, it was a scenario of one cell nibbling on another over time, rather than gobbling it up in one mouthful.
Where do the scientists get this idea? Lokiarchaeum.
Nicknamed "Loki," the microbe discovered last year may have evolved from the same lineage that gave rise to those first eukaryotes. Loki is an archaeon, but actually contains many of the same genes found in eukaryotes, but never before found outside them.
Scientists have determined that eukaryotes have two or three different evolutionary signatures in their genomes, says Dr. Dacks: "One of a bacterial origin, one of an archaeal origin, and then a whole category of genes where the signal is just not clear" or has been changed significantly in the eukaryotes.
And while scientists have known where to look for ancestors among bacteria, they didn't know where to look among the less well-known archaea.
The idea is that Loki is also descended from the eukaryote-ancestors, or at least was similar to them.
And those ancestors may have been archaea with genes that "allow them in a sense to be primed to acquire new characteristics once they encounter the complementary bacterial genes," Dey explains.
So studying Loki could help scientists tease out details about those ancestral cells.
"Now you can start teasing apart which parts of eukaryotes came from an archaeal contributor and which parts came from a bacterial contributor, and you just start closing down that gap," Dacks says. "What we get now are rigorous analyses that can speak to the areas of uncertainty and narrow down the possibilities" of how eukaryotes arose.
But there's one problem.
Scientists have yet to actually see Loki cells. The organism was discovered by scientists who were sequencing DNA from sediments bored out of the bottom of the Arctic Ocean.
That team of scientists used new technology to sequence all the genetic material in those sediments and sort out different sequences to identify species.
"It's like if you went into a toy store and you got every jigsaw puzzle, dumped all the pieces out, and then you used a computer to reassemble those jigsaw puzzles, including those jigsaw puzzles that you didn't even know were there," Dacks says, explaining the new technology.
Scientists hope to get their hands on some actual Loki cells, or some cells from its relatives.
Dey, who was not involved in sequencing the genome himself, but pored over it for this new paper, says that, "Based on our analysis of this gene content ... without the bacterial contribution, we predict that it won't look like a complex eukaryote inside."
But, Baum says their predictions could be wrong. "Nature often surprises you," he says. "And that's the fun thing about science."